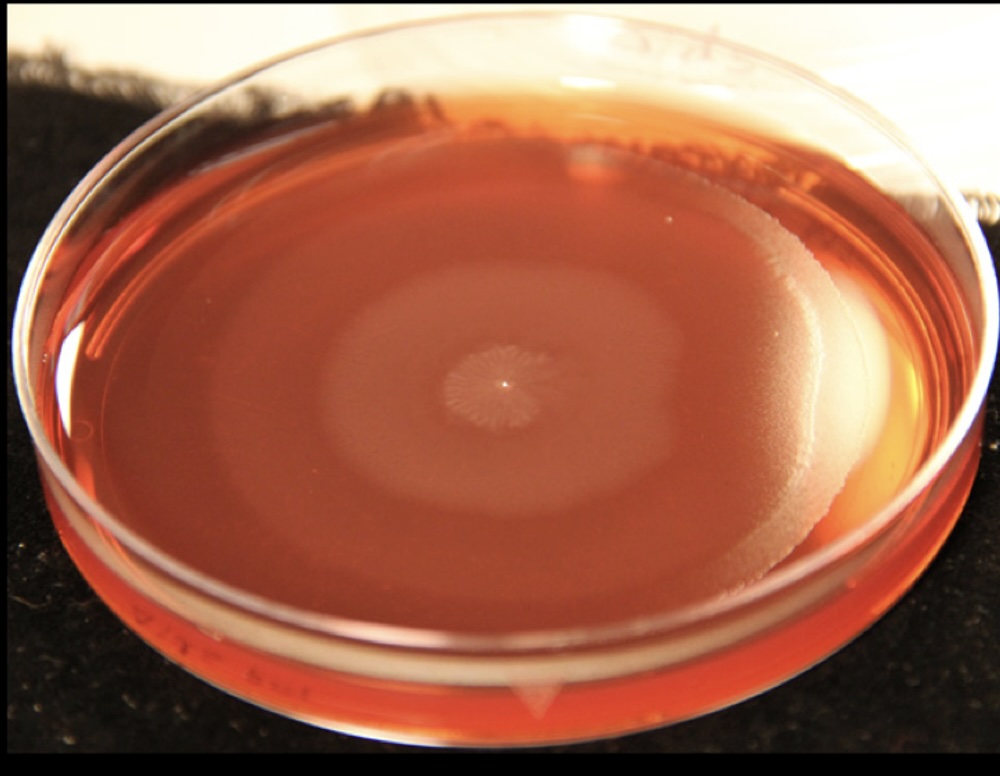
Proteus mirabilis, a strain of bacteria that naturally occurs in the gut, is one of the leading causes of antibiotic-resistant urinary tract infections in catherized patients. When a P. mirabilis colony begins migrating (or "swarming"), both the geometry of the individual bacteria and the shape of the overall colony undergo drastic changes. Individual cells elongate, align, and move, aggregating into a variety of higher-level structures as the leading edge of the swarm moves forward. Formal characterization of these geometric changes is key to assessing experimental results and understanding the swarming behavior.

The concentric rings in the image above, which shows a P. mirabilis colony that has grown outwards from a central inoculation point, result from the cyclical nature of the swarming behavior: ~3 hours of doubling, followed by ~2 hours of active movement, then roughly an hour of cell division before the cycle repeats. Details: cells from strain BB2000 were incubated at 37°C on a CM55 blood agar plate containing a dye for better contrast. Photographs were taken by Andrew Fields with a Canon SLR at an oblique angle. Owner: Gibbs Lab @ UC Berkeley.
During this cyclical process, the leading edge of the swarm develops a complex, filigreed structure, as shown in these closeups, taken two hours apart during an eight-hour experimental run:
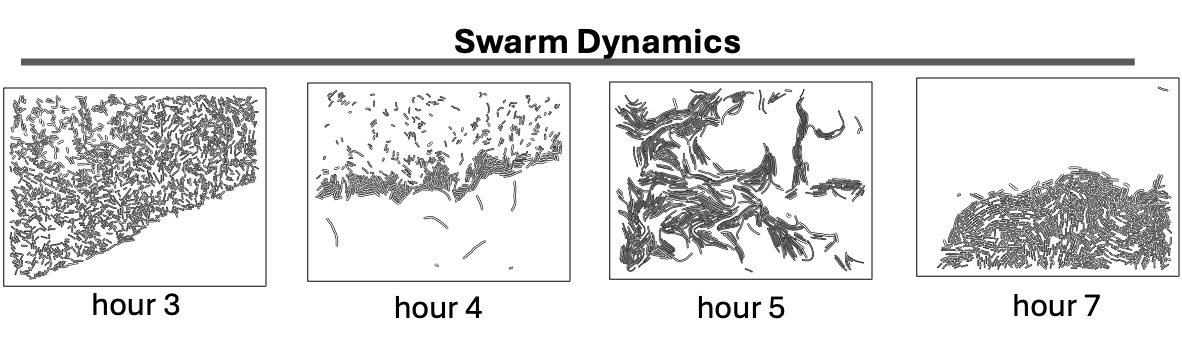
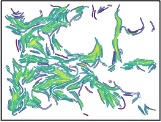
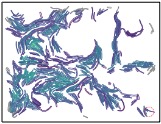

In the field of microbiology, there is a pressing need for language and tools to analyze individual interactions within a bacterial swarm, particularly those that are surface-bound. Our initial results are promising, as they suggest that we can successfully define key aspects of cell behaviors in collective migration. These include cell adjacency, cell-cell networks, geospatial density maps, and geometrical descriptions of colony edges, all at a remarkably high resolution. The potential impact of this research is significant, as it forms the foundation for future studies that could lead to a deeper understanding of bacterial behavior and pave the way for new strategies to disrupt intercellular interactions in mutant strains.
Liz & Karine would also like to thank the Packard Foundation for helping nucleate this project.
